RFP lab
Purpose
To express red fluorescent protein from jellyfish in bacteria. This expirement would also help us learn about genetic engineering.
Materials
Lab 2a
Materials can be found in Amgen lab manual part 2a.
Lab 4a
Materials can be found in Amgen lab manual part 4a.
Lab 5a
Materials can be found in Amgen lab manual part 5a.
Lab 6
Materials can be found in Amgen lab manual part 6.
Experimental Overview
Lab 2a - We verified that we had the correct plasma by using a restriction digest. We cut the plasmid with BamHI and Hind III.
Lab 4a - We verified the plasmid digest by electrophoresis.
Lab 5a - Then we transformed the bacteria with a recombinant plasmid.
Lab 6 - The we purified the RFP using chromotography.
To express red fluorescent protein from jellyfish in bacteria. This expirement would also help us learn about genetic engineering.
Materials
Lab 2a
Materials can be found in Amgen lab manual part 2a.
Lab 4a
Materials can be found in Amgen lab manual part 4a.
Lab 5a
Materials can be found in Amgen lab manual part 5a.
Lab 6
Materials can be found in Amgen lab manual part 6.
Experimental Overview
Lab 2a - We verified that we had the correct plasma by using a restriction digest. We cut the plasmid with BamHI and Hind III.
Lab 4a - We verified the plasmid digest by electrophoresis.
Lab 5a - Then we transformed the bacteria with a recombinant plasmid.
Lab 6 - The we purified the RFP using chromotography.
Data results
Before the 2a Lab:
1. Two fragments are produced. The two fragments are RFP with pBAD and Ara-C with ori with Amp-R.. The RFP plus pBAD is 807 BP, and Ara-C, ori, and Amp-R is 4495 BP.
2. The RFP gene and Ara-Care are the needed components in the plasmid .
3. The selectable marker allows only the desired bacteria to grow. The selectable marker separates the bacteria from the desired gene.
2a Questions:
1. Ori = origin of replication; RFP = red fluorescent protein (gene of interest); Amp-R = selectable marker; Ara-C = binds to promoter so we get transcription of gene of interest.
2. Restriction enzymes are a defense mechanism. They are there to cut up foreign bodies.
3. Bacteria would retain a gene that give them resistance to antibiotics to protect themselves from disease.
4. The central dogma in each organism is the same.
5. Grow Kan and Amp bacteria in a petri dish. Put half of the mixed bacteria into each separate petri dish petri dish and wait a day. The bacteria should have died and then you should be able to separate them apart.
4a Questions:
1. The purpose of growing bacteria that resists ampicillin is to make sure that the desired cell lives.
2. If the bacterial cells are withheld of arabinose, then the pARA-R plasmid will not turn on the promoter. Then the protein will not turn red.
3. In the LB plate, both the P- and P+ will grow. In the LB/Amp plate, no P- will grow, but P+ will. In the LB/ Amp/Ara plate, there will be little bacteria growth.
5a Questions:
1. Our predictions touched the actual results. The LB plate had limited growth. The LB/Amp and LB/Amp/Ara plate matched our prediction.
2. There were no red colonies visible either due to temperature or not enough time in the incubator.
3. The LB-amp-ara plate prevents transcription to occur, which allows RFP to be expressed. The LB-amp does not have Ara.
4. The multiple copies are important because it gives a greater chance of the promoter being turned on.
5. The RFP gene is expressed as a trait through transcription (central dogma).
6a Questions:
1. The red fluorescent protein has red cells and thats is why it can be separated.
2. The pellet is pink. The supernatant is clear liquid.
6b Questions:
1. Binding Buffer (BB): causes amino acid and protein bind to the resin beads.
Wash Buffer (WB): removes loose proteins that are not bound to the resin beads.
Elution Buffer (EB): takes off protein off resin beads.
Column Equilibration Buffer (CEB): stores resin beads.
2. The supernatant was more pink than clear. However, the pellet was a little darker pink than the supernatant.
1. Two fragments are produced. The two fragments are RFP with pBAD and Ara-C with ori with Amp-R.. The RFP plus pBAD is 807 BP, and Ara-C, ori, and Amp-R is 4495 BP.
2. The RFP gene and Ara-Care are the needed components in the plasmid .
3. The selectable marker allows only the desired bacteria to grow. The selectable marker separates the bacteria from the desired gene.
2a Questions:
1. Ori = origin of replication; RFP = red fluorescent protein (gene of interest); Amp-R = selectable marker; Ara-C = binds to promoter so we get transcription of gene of interest.
2. Restriction enzymes are a defense mechanism. They are there to cut up foreign bodies.
3. Bacteria would retain a gene that give them resistance to antibiotics to protect themselves from disease.
4. The central dogma in each organism is the same.
5. Grow Kan and Amp bacteria in a petri dish. Put half of the mixed bacteria into each separate petri dish petri dish and wait a day. The bacteria should have died and then you should be able to separate them apart.
4a Questions:
1. The purpose of growing bacteria that resists ampicillin is to make sure that the desired cell lives.
2. If the bacterial cells are withheld of arabinose, then the pARA-R plasmid will not turn on the promoter. Then the protein will not turn red.
3. In the LB plate, both the P- and P+ will grow. In the LB/Amp plate, no P- will grow, but P+ will. In the LB/ Amp/Ara plate, there will be little bacteria growth.
5a Questions:
1. Our predictions touched the actual results. The LB plate had limited growth. The LB/Amp and LB/Amp/Ara plate matched our prediction.
2. There were no red colonies visible either due to temperature or not enough time in the incubator.
3. The LB-amp-ara plate prevents transcription to occur, which allows RFP to be expressed. The LB-amp does not have Ara.
4. The multiple copies are important because it gives a greater chance of the promoter being turned on.
5. The RFP gene is expressed as a trait through transcription (central dogma).
6a Questions:
1. The red fluorescent protein has red cells and thats is why it can be separated.
2. The pellet is pink. The supernatant is clear liquid.
6b Questions:
1. Binding Buffer (BB): causes amino acid and protein bind to the resin beads.
Wash Buffer (WB): removes loose proteins that are not bound to the resin beads.
Elution Buffer (EB): takes off protein off resin beads.
Column Equilibration Buffer (CEB): stores resin beads.
2. The supernatant was more pink than clear. However, the pellet was a little darker pink than the supernatant.
data analysis
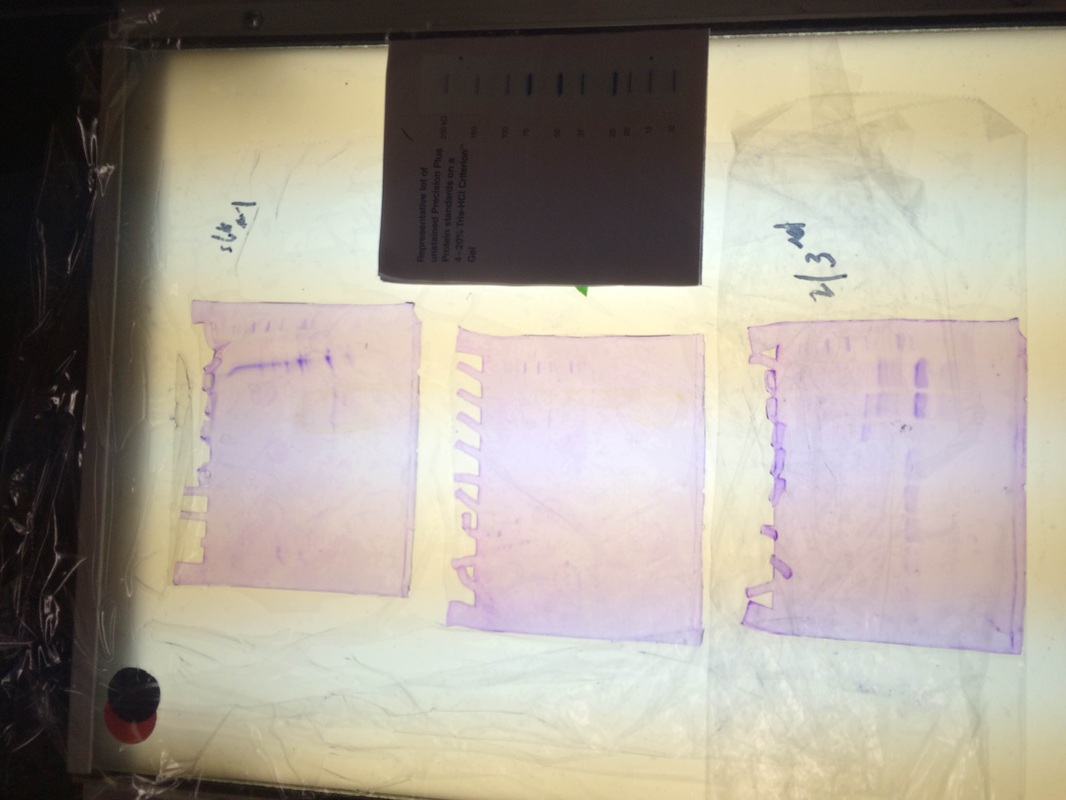
Our column for loading did not show up, but our R+ and R- both showed perfectly.
We extracted the RFP, then ran it through a gel.
Our column was on the first one and it showed the purple line. However, our gel had some problems, as you can tell from the weird formation, but when we looked at second and third periods gel we saw that some of their groups had quite good RFP extraction. Their bands were thick and in the proper place. (According to the chart above).
reflection
1. What did you like/find interesting?
I liked being able to see the pink DNA and I found that you are able to pick what bacteria can grow on what plate. I am also interested in what different types of bacteria look like.
2. How did you and your partner collaborate?
My group was Sawyer, Jonathan, and I. We worked well together and at first ti was confusing. Slowly over time I got what we were doing and became very interested in the lab. This is ones of the better more efficient groups I have been in. Maybe because its smaller.
3. What would you do differently next time?
Next time, I would of had the lab be more simple. It was a little complex, and it moved fast in some places, but I got understood what was going on. I would of got it faster if the instructions were more simple.
I liked being able to see the pink DNA and I found that you are able to pick what bacteria can grow on what plate. I am also interested in what different types of bacteria look like.
2. How did you and your partner collaborate?
My group was Sawyer, Jonathan, and I. We worked well together and at first ti was confusing. Slowly over time I got what we were doing and became very interested in the lab. This is ones of the better more efficient groups I have been in. Maybe because its smaller.
3. What would you do differently next time?
Next time, I would of had the lab be more simple. It was a little complex, and it moved fast in some places, but I got understood what was going on. I would of got it faster if the instructions were more simple.